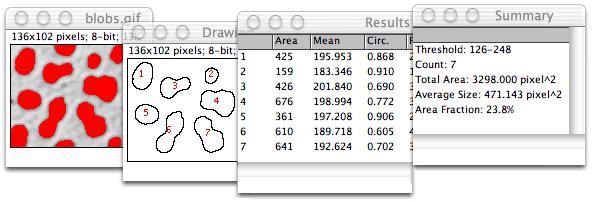 
However, they depend on prior object detection and segmentation. Object-based methods are more intuitive to interpret, as they directly work with locations of objects. This, however, leads to a loss of information as nearby molecule detections fuse into one blob in the rendered image. Pixel-based methods are hence not directly applicable, unless one first renders a synthetic image from the observed point locations. Moreover, in some observations, like those from Photo-Activation Localization Microscopy (PALM) and STochastic Optical Reconstruction Microscopy (STORM), it is not obvious what constitutes an image, as these methods provide locations and localization uncertainties of individual molecules. They are also sensitive to blurring and noise in the image. While pixel-based measures are easy to compute, they are difficult to interpret. Object-based methods first detect and delineate the objects of interest in the images and then quantify their degree of co-localization using an overlap measure. Pixel-based methods typically use a correlation measure between the pixel intensities in different images in order to quantify the degree of overlap or co-localization between the object distributions represented in the images. This includes pixel-based and object-based co-localization analysis methods. 
īiology has long relied on co-localization analysis in order to quantify the spatial distribution of one set of objects with respect to another one. Significant deviations from spatial randomness are indicative of interactions (of some sort) between the objects, as formalized in the framework of spatial interaction analysis. This for example allows testing whether the distribution of a set of objects is significantly different from random, or whether the objects in one set are distributed independently of the objects in another set. The mathematical framework of spatial statistics allows quantifying and analyzing such patterns, comparing them with each other, and performing statistical tests on them. This ranges from spatial patterns of sub-cellular structures or proteins, to the spatial patterns formed by cells in tissues, to spatial patterns of organisms in ecosystems. The spatial arrangement of objects relative to each other is a rich source of phenotypic information. 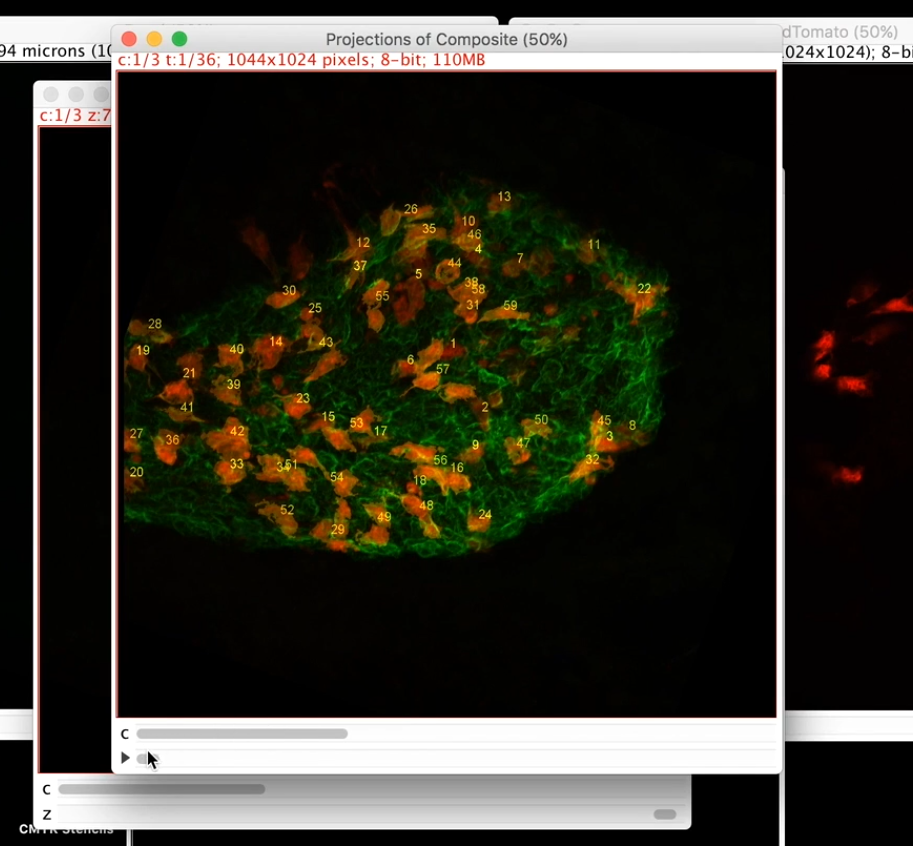
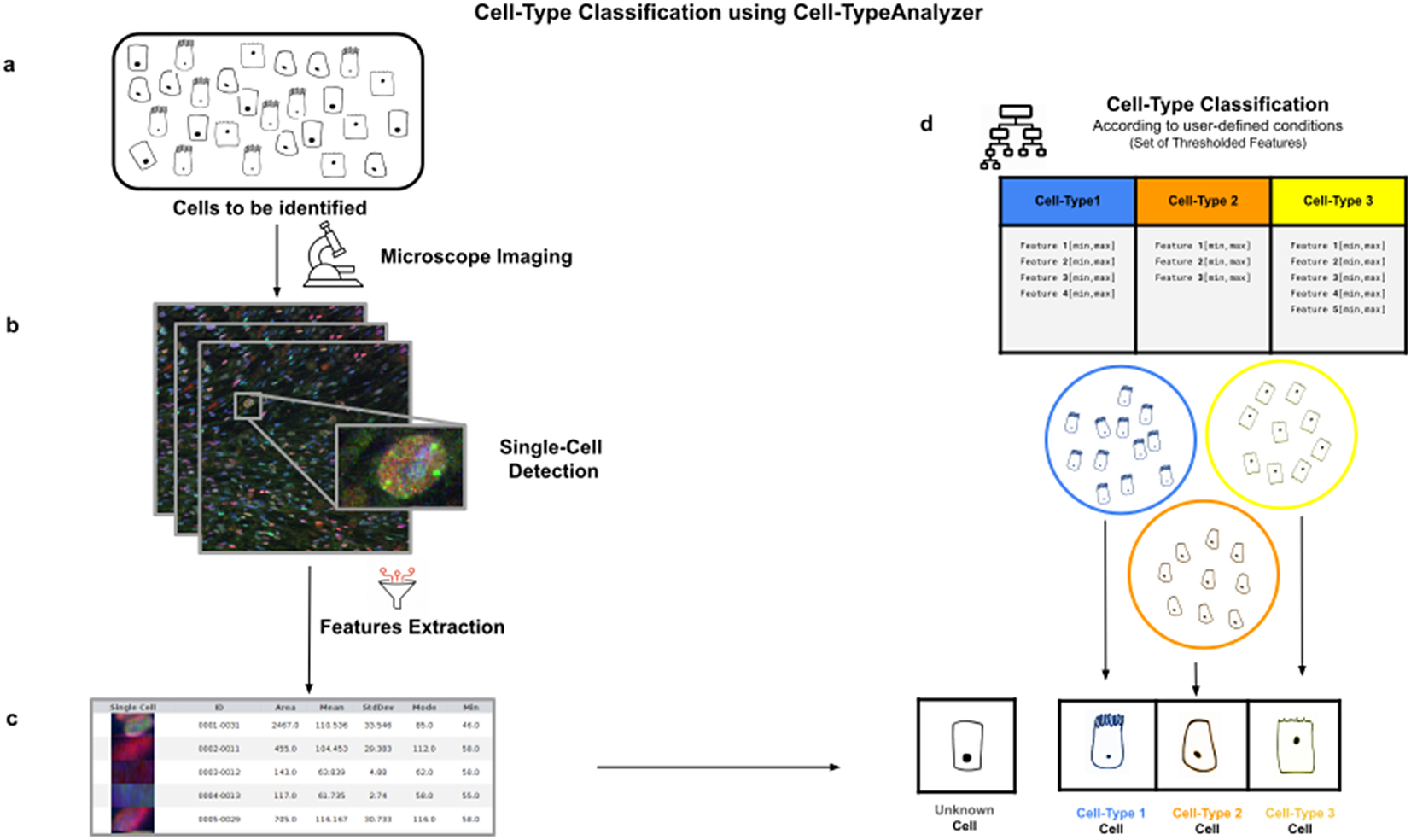
We present a software plugin to analyze and quantify spatial patterns of objects in images using the free open-source image-processing platform ImageJ or its distribution Fiji. The presented showcases illustrate the usage of the software. The present software greatly simplifies spatial interaction analysis for point patterns, and makes it available to the large user community of ImageJ and Fiji. We benchmark and demonstrate the present software using examples from confocal and PALM single-molecule microscopy. The plugin detects objects in images, infers the interaction potential that is most likely to explain the observed pattern, and provides statistical tests for whether an inferred interaction is significant given the number of objects detected in the images and the size of the space within which they can distribute. We present an ImageJ/Fiji plugin that implements the complete workflow of spatial pattern and interaction analysis for spot-like objects. However, no user-friendly software for this type of analysis was available so far. This can be used to generalize co-localization analysis to spatial interaction analysis. Spatial correlations in their distributions are indicative of interactions and can be modeled by an effective interaction potential acting between the points of the two sets. If they do not “feel” each other’s presence, their spatial distributions are expected to be independent of one another. The relative arrangement of a set of objects with respect to another set of objects contains information about potential interactions between the two sets of objects. Analyzing spatial distributions of objects in images is a fundamental task in many biological studies.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
